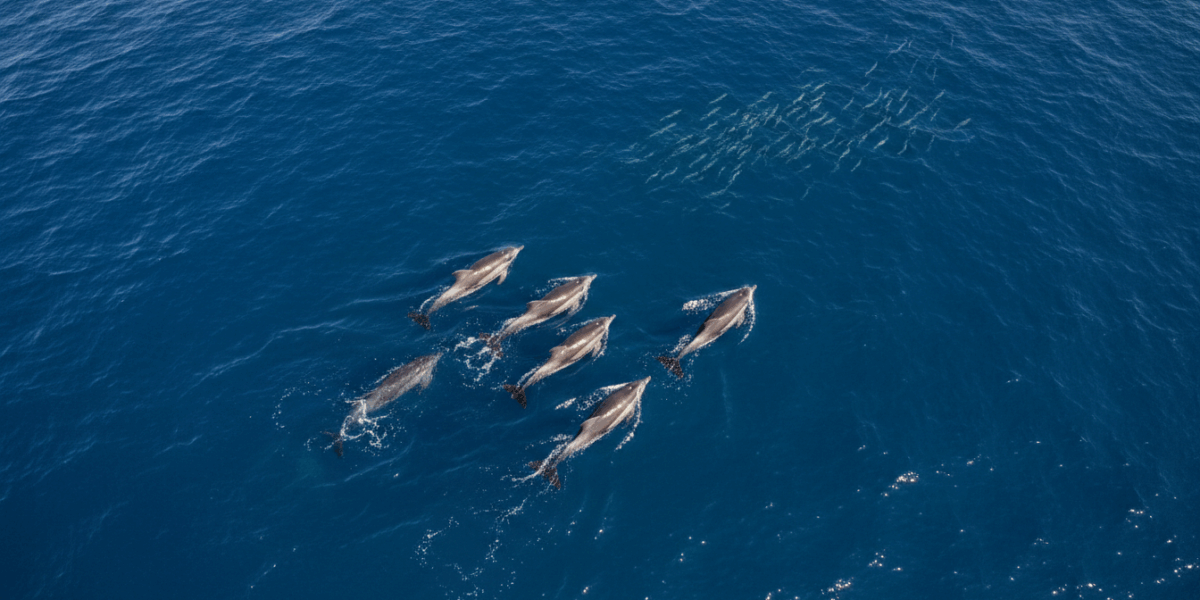Why This Matters
Dolphins are iconic marine species in Aotearoa New Zealand and globally. They play key ecological roles as apex or mesopredators, structuring marine food webs and acting as indicators of ocean health. New Zealand is home to several endemic and threatened species, including Hector’s and Māui dolphins, which face increasing pressures from fisheries interactions, habitat degradation, climate-driven shifts in prey availability, pollution, and cumulative human impacts.
Understanding dolphin ecology, particularly diet and health status, is critical for informed conservation and adaptive management in a rapidly changing marine environment. Detecting shifts in prey use over time and space helps reveal how dolphins respond to environmental variability, ecosystem change, and anthropogenic stressors.
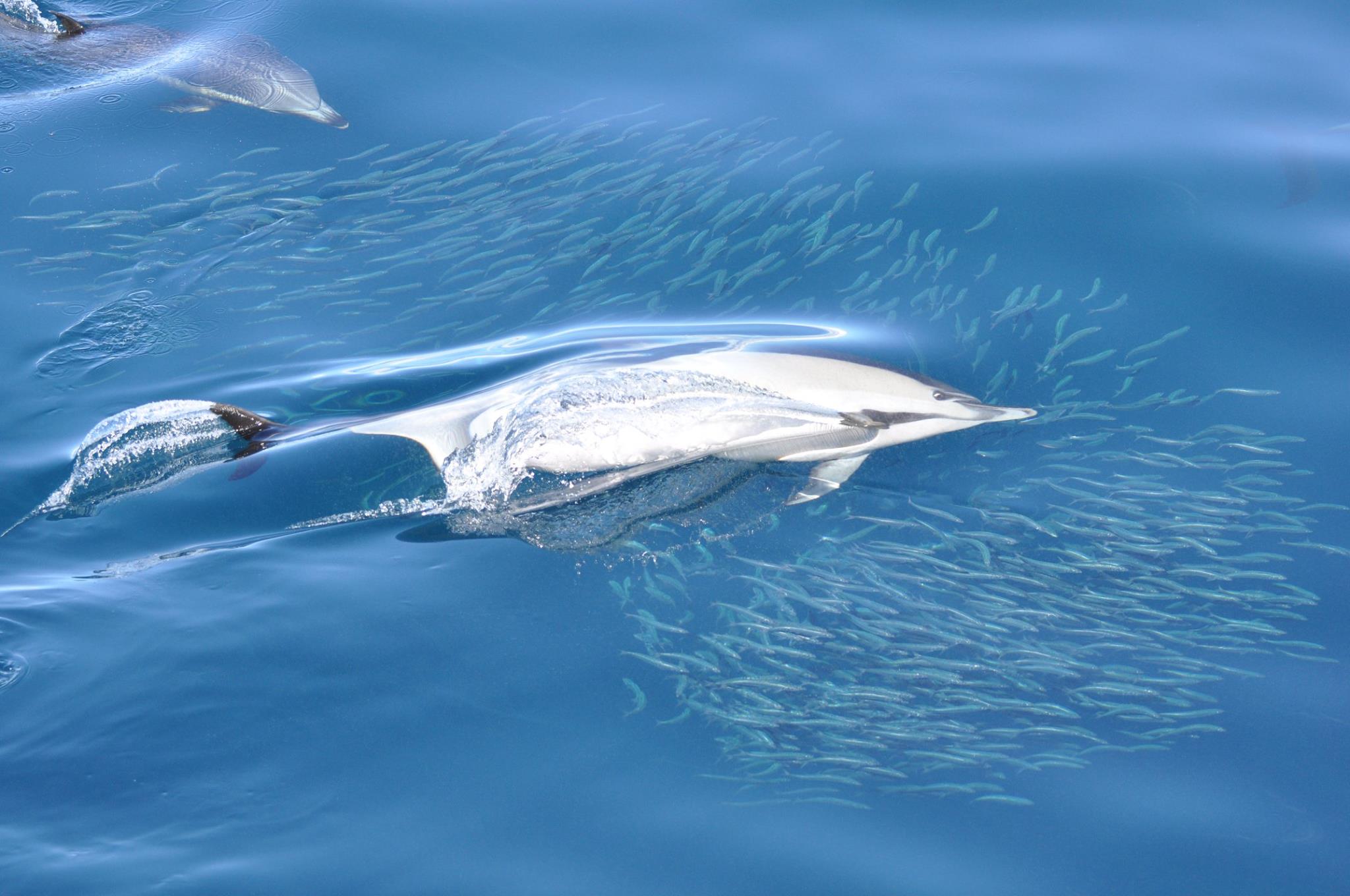

Dolphins feeding at sea. Photo credit: Auckland Whale and Dolphin Safari (AWADS)
Moving Beyond Traditional Diet Analysis
Diet studies in stranded dolphins have traditionally relied on hard part analysis during necropsy, identifying prey remains such as otoliths and cephalopod beaks. While valuable, this approach can underestimate soft-bodied prey, highly digested material, or taxa that leave no identifiable remains.
Environmental DNA and metabarcoding offer a powerful complementary approach. By analysing DNA extracted from stomach contents and gut material, it is possible to detect a broader spectrum of prey species, including those missed by conventional methods. This provides higher taxonomic resolution and a more comprehensive view of trophic interactions.
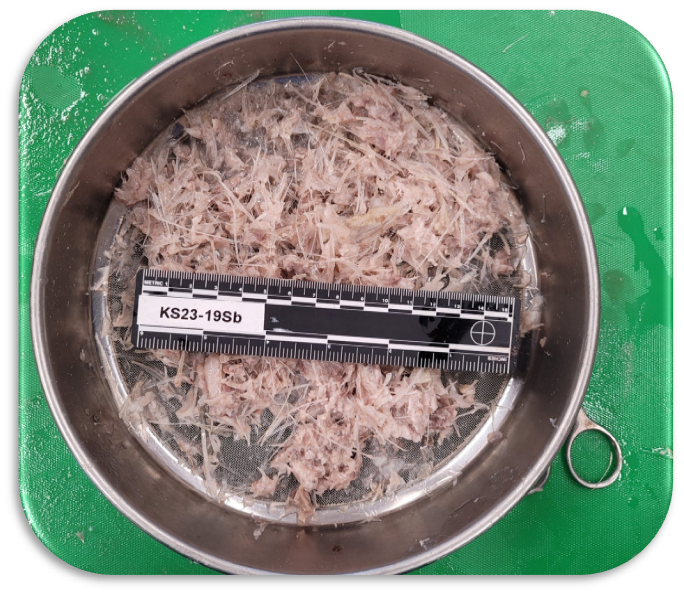


Gut contents sample (left) and homogenized samples for eDNA analyses (right). Photo credit: Cetacean Ecology Research Group (CERG)
Sequench Contribution
Sequench is excited to contribute molecular expertise to ongoing research led by staff and students at the Cetacean Ecology Research Group (CERG) at Massey University, Auckland. One project supported by Auckland Whale and Dolphin Safari investigates temporal and spatial patterns of diet shifts in New Zealand dolphin species using high-throughput eDNA metabarcoding.
In addition to prey identification, the research further explores the gut microbiome of dolphins to better understand associations between microbiome diversity, taxonomic composition, and host health status. Linking dietary information with gut microbial structure provides new insight into physiological condition, ecological adaptation, and potential stress responses.
This collaboration strengthens the evidence base for dolphin conservation in New Zealand waters and demonstrates how molecular tools can enhance marine wildlife research, enabling more precise and forward-looking management strategies.
Join Us in Advancing Research
Do you have an innovative idea or a research question? We’re always eager to collaborate with passionate individuals and organizations. Contact us today to explore how we can work together to bring your vision to life and make a lasting impact on our planet.